
Mapping Algorithms - How They all Arrived
We have been polishing up the tutorials on alignment programs and wanted to share few ideas.
This page shows an enormous explosion of the number of available mappers over the last five years.
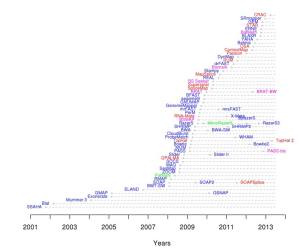
When you go into their codes and algorithms (we have a section covering most popular programs), you see that they mostly fall into handful of broad categories. Those categories are best presented in Mark Chaisson’s informative figure from BLASR paper.
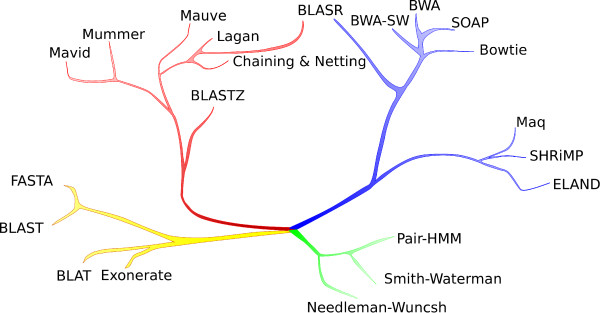
It is also important to note that time-wise, nodes in each branch appeared almost together and their development was as much the result of needs from biologists as the non-biological development in computer science or electrical engineering. For example, MUMmer, AVID, etc. were all released after 1998, when the suffix array sub-field of computer science matured enough. SSAHA utilized the availability of large amount of RAM in a box, not unlike what SOAPdenovo assembler did 8-9 years later at a bigger scale to assemble the panda genome. Even though Burrows Wheeler transform was published in 1994, it could not be used in search programs until Paolo Ferragina and Giovanni Manzini showed a way in their 2000 and 2005 seminal papers.
Hopefully, you will find all those discussions useful in our tutorial section. We will continue to improve them.