
What Will Replace GATK in the BALSA Age?
Broad Institute and HKU/BGI are two major contributors for algorithms in the next-gen era. Our earlier commentary “BALSA: Integrated secondary analysis for whole-genome and whole-exome sequencing, accelerated by GPU” may have given the impression that BALSA increases speed of variant calling by many times (‘BALSA finished in 5.49 hours, whereas ISAAC finished in 11.92 hours, and GATK coupled with BWAaln, BWAmem and SOAP3-?dp in 88.00, 48.68 and 46.27 hours, respectively.’) and is an obvious replacement for current programs. That is assuming that researchers at Broad and elsewhere are not trying to improve existing pipelines, which is not true.
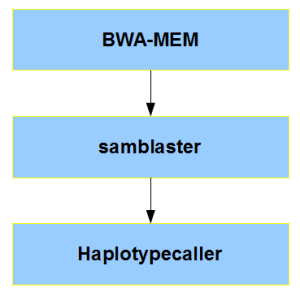
Here is a proposed modification of the
original GATK pipeline that will potentially lead to faster execution (h/t: hengli), although we do not have any benchmarks for comparison. Maybe, those who like to be on the cutting-edge of things, can try to compare BALSA with the following pipeline and let the community know about the increase in speed.
BWAmem is available here. It comes as one function of the BWA pipeline.
BWA is a software package for mapping low-divergent sequences against a large reference genome, such as the human genome. It consists of three algorithms: BWA-backtrack, BWA-SW and BWA-MEM. The first algorithm is designed for Illumina sequence reads up to 100bp, while the rest two for longer sequences ranged from 70bp to 1Mbp. BWA-MEM and BWA-SW share similar features such as long-read support and split alignment, but BWA-MEM, which is the latest, is generally recommended for high-quality queries as it is faster and more accurate. BWA-MEM also has better performance than BWA-backtrack for 70-100bp Illumina reads.
-—————————–
Samblaster is available here.
Samblaster is a fast and flexible program for marking duplicates in read-id sorted paired-end SAM files. It can also optionally output discordant read pairs and/or split read mappings to separate SAM files, and/or unmapped/clipped reads to a separate FASTQ file.
When marking duplicates, samblaster will require approximately 20MB of memory per 1M read pairs.
The samblaster paper is worth reading.
SAMBLASTER: fast duplicate marking and structural variant read extraction
-—————————–
Haplotypecaller is available here . It is part of the GATK pipeline.
Call SNPs and indels simultaneously via local de-novo assembly of haplotypes in an active region.
Please feel free to suggest other replacements for the above programs.