
How Does the Unix Diff Algorithm Work?
A link from stackoverflow says -
The basic algorithm is described in “An O(ND) Difference Algorithm and its Variations”, Eugene W. Myers, ‘Algorithmica’ Vol. 1 No. 2, 1986, pp. 251-266; and in “A File Comparison Program”, Webb Miller and Eugene W. Myers, ‘Software –Practice and Experience’ Vol. 15 No. 11, 1985, pp. 1025-1040. The algorithm was independently discovered as described in “Algorithms for Approximate String Matching”, E. Ukkonen, `Information and Control’ Vol. 64, 1985, pp. 100-118.
We checked Myers’ original paper and found an elegant graph-based greedy algorithm.
The problems of finding a longest common subsequence of two sequences A and B and a shortest edit script for transforming A into B have long been known to be dual problems. In this paper, they are shown to be equivalent to finding a shortest/longest path in an edit graph. Using this perspective, a simple O(ND) time and space algorithm is developed where N is the sum of the lengths of A and B and D is the size of the minimum edit script for A and B. The algorithm performs well when differences are small (sequences are similar) and is consequently fast in typical applications. The algorithm is shown to have O(N+D2 ) expected-time performance under a basic stochastic model. A refinement of the algorithm requires only O(N) space, and the use of suffix trees leads to an O(NlgN +D2 ) time variation.
The paper is well-written, but for those looking for additional explanation may find the following website helpful -
Investigating Myers’ diff algorithm: Part 1 of 2
Investigating Myers’ Diff Algorithm: Part 2 of 2
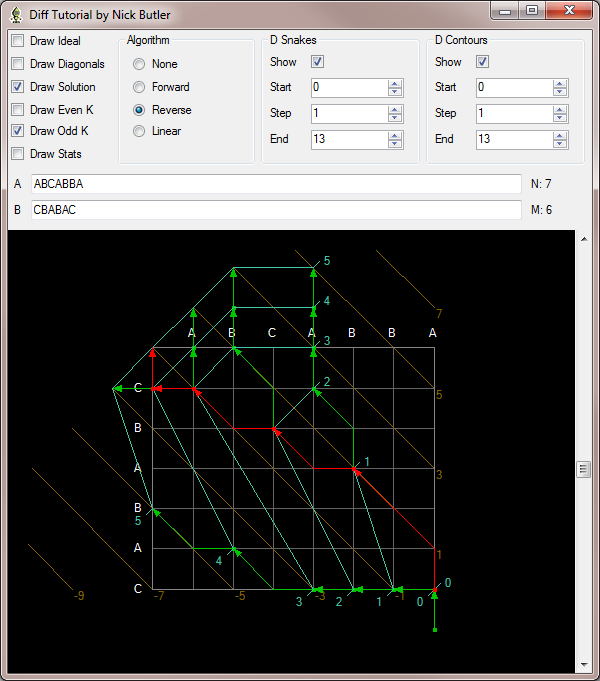
In bioinformatics sequence comparison, can we start doing ‘diff’ between sequences instead of going through other elaborate search programs? Such as taking 1 million long pacbio reads and doing ‘diff’ between pairs to see whether they are closely related?
More on that in a later commentary.