
An Email Interview with Evolutionary Developmental Biologists Isabelle Peter and Eric Davidson
I found the book “Genomic Control Process - Development and Evolution” by Professors Isabelle Peter and Eric Davidson very informative and thought-provoking, and contacted its authors to check whether they would be interested in publishing an interview to be shared with our readers. I am greatly honored that they agreed about not one, but two interviews. The first one (see below) is email- based, where they replied to a set of questions regarding their book sent by me. A video interview will follow next month.

Professor Davidson had been the source of inspiration for my work for many years. I first met him about a decade back, when I drove to Caltech to discuss about the possibility of using tiling arrays for the sea urchin genome project. I prepared a number of slides to make a convincing case, and was pleasantly surprised to find that he already knew far more about the technology than what I had in my slides. Then three years later (2007), Davidson was the first to warn me that the arrays would go away and be replaced by Illumina short-read sequencing.
The truly visionary aspect of his group’s research is in taking advantage of such technological advances to answer how the genomic features are linked to biological phenomena. In a series of recent papers, Dr. Peter and Dr. Davidson reported about making major empirical and conceptual breakthroughs and their book resulted from that collaborative work. However, the book is not limited to their own work, but provides novel synthesis for a large spectrum of ‘evo- devo’ in animals.
The questions and answers are enclosed below.
-——————————————————————


1. Would you please give us a brief summary of what the book is about?
Peter/Davidson: This book is about the mechanisms by which the linear genomic DNA sequence encodes major processes of animal life, the most demanding of which is development. In this book we have treated a vast array of biological phenomena, by organizing them around a framework concept, the concept of gene regulatory networks. These networks are systems of interacting regulatory genes, which ultimately determine the spatial and temporal expression of all genes in the developing animal. The same arguments inform what has long been regarded as the most fundamental problem in biology, which is how evolution works. In our book we have dealt with mechanisms of gene regulation in animal cells, the mechanisms of development in many different contexts, and the mechanisms by which animal body plans have diverged in deep time. What this book is really about is how the fundamental principles of genomic information use give us an explanatory framework and enables us to see any aspects of animal life through that same lens.
2. What aspects of animal life is this book focused on?
Peter/Davidson: There are four large general areas for which this book provides an in-depth interpretation. The first of these areas deals with the molecular biology of gene regulatory networks and evaluates how sequence-specific gene regulation is achieved in animal cells. Several regulatory mechanisms, such as transcriptional regulation, non-coding RNAs and epigenetic histone modifications, are discussed, all of which ultimately depend on the sequence recognition function of transcription factors. Second, we consider the function of gene regulatory networks in controlling the processes of development from egg to embryo, from embryo to body part, and from body part to cell types. Third we consider the functional import of the structure of developmental networks. Three very important aspects that we discuss in depth are the deep hierarchy of these networks; their modular structure, in that they are composed of what seems to be a finite set of subcircuit architectures; and their primary function in organizing an increasingly complex landscape of spatial expression domains. We also compare diverse computational approaches to network function. Fourth, we consider evolutionary process as the outcome of change and stasis since the beginning of the Cambrian, in the structure of developmental networks encoded in animal genomes.
3. Are there areas outside development and evolution per se that your book impacts?
Peter/Davidson: The book has broad implications for many contiguous areas of bioscience, in addition to development and evolution. These include the mechanisms of mobilization of effector genes, often of medical significance, that do all the work of differentiated cells, and many kinds of physiological response. All of these ultimately require understanding of genomic control mechanisms, a subject that converges on the basic theoretical conceptions of this book. This casts in a new light the definition of functional genomics, in that the functional genome consists of the components of gene regulatory networks, largely those encoding developmental process.
4. What audience is the book addressed to?
Peter/Davidson: We have written the book with a broad audience in mind, essentially anyone who has a background in science or engineering and some basic level of familiarity with molecular biology. Although the arguments of the book arrive at the growing point of knowledge, concepts and processes are all defined from the ground up, so that anyone who cares can follow. We have deliberately sought to write this book on two simultaneous parallel planes: readers concerned with the major ideas and concepts can enjoy these without emerging into biological detail, while those concerned with experimental evidence will find ample representation of this level of reality, particularly in the figures.
5. Is this book comparable to a textbook or an extended review of a large field of developmental biology?
Peter/Davidson: This book is not a review of received wisdom, but a novel synthesis informed by diverse areas of biology. We feel that a synthesis is particularly important in our time, when there is a general sense that biology has become an overwhelming mass of context-specific details and data that make the extraction of broad and incisive mechanistic principles utterly impossible. Although in a conventional sense this book is not a textbook, we have used it as a basis for a new mode of concept-oriented teaching.
6. What is the rationale for the novel synthesis you mention?
Peter/Davidson: Many aspects of biology depend directly on information carried in the genome, and the control system for genomic function thus represents something like a common denominator of biology. Indeed, genomic control logic provides a common mechanistic basis for the most disparate processes, ranging from specific cell function to embryogenesis to making of the body plan to body plan evolution. Being able to think about these processes in common mechanistic terms offers an immensely powerful tool to compare and relate traditionally distinct aspects of animal biology.
7. In Chapter 6 you present your new computational Boolean treatment of developmental gene regulatory networks: what is its significance?
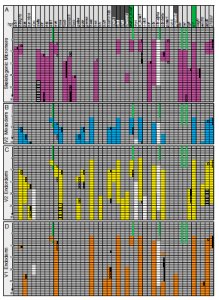
Peter/Davidson: This model was a successful attempt to utilize the experimentally defined regulatory interactions of a developmental gene regulatory network to reconstruct in silico developmental gene expression in time and space, with very considerable accuracy. The result was a proof of principle that causality in gene expression, i.e. the basis of development, lies in encoded gene regulatory networks, and that such networks can be experimentally solved. This work also shows that experimentally studying gene regulatory networks represents the most important pathway towards understanding large areas of biological process. Thus the Boolean model unlocks a major conceptual road to the future.
-—————————————————————
-—————————————————————
Homolog.us blog thanks the publisher (Elsevier) for sharing a chapter with our readers and for sponsoring our work. If you are purchasing at the Elsevier online bookstore, please use the code BIOMED315 to get a 30% discount on the price.