
Evolutionary Biologists to Follow - Sally Leys and Casey Dunn
If there is one area of evolutionary biology benefiting greatly from high- throughput sequencing, that is the study of ‘primitive’ or ‘lower’ animals. The animals like sponges were traditionally avoided in the era of model organisms, because what can one really learn about humans based on brainless, heartless and limbless ‘simple’ animals? Surprisingly, high-throughput sequencing found those animals not to be as simple as previously thought. It is also worth pointing out the roles of two researchers (Sally Leys and Casey Dunn), who are truly pushing the boundaries by taking advantage of new sequencing techniques and asking interesting questions.
-——————————————–
Six major steps in animal evolution: are we derived sponge larvae?
The importance of sponges and choanoflagellates in understanding evolution were highlighted in a 2009 review paper by Claud Nielsen of Natural History Museum of Denmark. Dr. Nielsen was the past president of International Society for Invertebrate Morphology, winner of Linnean Medal and the author of widely cited book ‘Animal Evolution: Interrelationships of the Living Phyla’. In the paper he wrote -
A review of the old and new literature on animal morphology/embryology and molecular studies has led me to the following scenario for the early evolution of the metazoans. The metazoan ancestor, choanoblastaea, was a pelagic sphere consisting of choanocytes. The evolution of multicellularity enabled division of labor between cells, and an advanced choanoblastaea consisted of choanocytes and nonfeeding cells. Polarity became established, and an adult, sessile stage developed. Choanocytes of the upper side became arranged in a groove with the cilia pumping water along the groove. Cells overarched the groove so that a choanocyte chamber was formed, establishing the body plan of an adult sponge; the pelagic larval stage was retained but became lecithotrophic. The sponges radiated into monophyletic Silicea, Calcarea, and Homoscleromorpha. Homoscleromorph larvae show cell layers resembling true, sealed epithelia. A homoscleromorph-like larva developed an archenteron, and the sealed epithelium made extracellular digestion possible in this isolated space. This larva became sexually mature, and the adult sponge-stage was abandoned in an extreme progenesis. This eumetazoan ancestor, gastraea, corresponds to Haeckel’s gastraea. Trichoplax represents this stage, but with the blastopore spread out so that the endoderm has become the underside of the creeping animal. Another lineage developed a nervous system; this neurogastraea is the ancestor of the Neuralia. Cnidarians have retained this organization, whereas the Triploblastica (Ctenophora+Bilateria), have developed the mesoderm. The bilaterians developed bilaterality in a primitive form in the Acoelomorpha and in an advanced form with tubular gut and long Hox cluster in the Eubilateria (Protostomia+Deuterostomia).
It is indicated that the major evolutionary steps are the result of suites of existing genes becoming co-opted into new networks that specify new structures.
The evolution of the eumetazoan ancestor from a progenetic homoscleromorph larva implies that we, as well as all the other eumetazoans, are derived sponge larvae.
-——————————————–
The hidden biology of sponges and ctenophores
In a review paper published in Trends in Ecology and Evolution, Casey Dunn, Sally Leys and Steven Haddock went a step further and challenged the notion of human-centric understanding of animals like sponges and ctenophores. The concept of ‘hidden biology’ is explained in Fig 1 of their paper and is shown below along with the extended caption (Box 1).
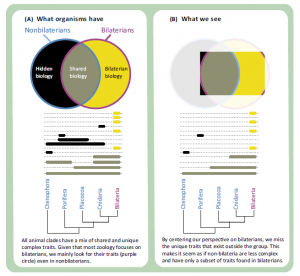
All living animals belong to one of five clades: Porifera, Ctenophora, Placozoa, Cnidaria, and Bilateria. To a first approximation, the study of zoology is the study of Bilateria. Humans and all the best-studied model animal species (mouse, Drosophila melanogaster, Caenorhabditis elegans, and others) are within Bilateria. All of the terrestrial animals, and most freshwater animals that humans regularly encounter are within Bilateria. Most known animal species are within Bilateria (in fact, most known species belong to a single bilaterian clade: Arthropoda). However, if we want to understand the full breadth of animal diversity and the earliest events in animal evolution, we need to study all animals, not just Bilateria.
To a large extent, the focus on the study of bilaterians is a resource allocation decisions: zoologists spend more time and money studying bilaterians than they do nonbilaterians because they comprise most living animal species, including ourselves and the animals we are most familiar with. However, this creates a problem: currently, we see most nonbilaterian biology through the filter of bilaterian biology (Figure I). All animal clades have a mix of unique traits and traits that are shared with other animals (Figure IA). It is easier to confirm previously known traits and functions than it is to describe new traits and functions, and most previous studies have been on bilaterians. In addition, many widely used tools and reagents have been optimized for Bilateria. This makes it easier to study the aspects of nonbilaterian biology that are similar to bilaterian biology (Figure IB, gray), than it is to study traits that are only found outside Bilateria (Figure IB, black). The candidate gene approach is a widespread example of this. However, just because it is easiest to study the subset of nonbilaterian biology that is shared with bilaterians does not mean that nonbilaterians only have a subset of bilaterian biology, or that bilaterians are more advanced than other animals. It just means that many of their unique features are currently unknown to us: a hidden biology (Figure I) that we have only the first glimpses of. This hidden biology includes novel structures and functions, facilitated by novel mechanisms, that are not found in bilaterian model species. It also includes novel mechanisms that underlie shared structures and functions. The problems of hidden biology also extend to nonmodel bilaterians, although it is more severe in nonbilaterians.
What do we miss by letting so much nonbilaterian biology stay hidden? At best, we miss out on some interesting biology, including unique morphology, developmental mechanisms, and physiology. At worst, we are systematically misled. Unfortunately, this is the case when it comes to understanding early animal evolution. It is tempting to mistake our biased perspective (Figure IB) for the actual distribution of traits (Figure IA), which gives the false impressions that nonbilaterians have only a subset of the traits found in Bilateria and, therefore, that they are lower or simpler. This in turn plays into the misconception that living animal diversity conforms to a linear aristotelian scala naturae, from lower to higher animals, and that animal evolution has proceeded by a step-wise accumulation of complex traits, such that the more distantly an animal is related to Bilateria, the more closely it resembles the most recent common ancestor of all animals. In reality, all living animal lineages have had the same amount of time to evolve since the most recent common ancestor of all animals, and all have gain and lost multiple traits. We need to understand the traits present in all animal groups, not just those that are present in Bilateria, if we are to understand early animal evolution.
-——————————————–
Sally Leys - Elements of a ‘nervous system’ in sponges

The claims of complexity of sponges, jellies and ctenophores do not appear far-fetched, if you go carefully through the papers of Dr. Sally Leys. For years, she collaborated with George O. Mackie and showed that jellyfish had central neural circuitry (check 2004 review ‘Central Neural Circuitry in the Jellyfish Aglantha’) and then moved her attention to sponges. Her work combines traditional electrical measurements and high-throughput sequencing. In the paper - ‘Evolutionary origins of sensation in metazoans: functional evidence for a new sensory organ in sponges’, she and co-authors showed that sponges have sensory cilium.
Background
One of the hallmarks of multicellular organisms is the ability of their cells to trigger responses to the environment in a coordinated manner. In recent years primary cilia have been shown to be present as antennae on almost all animal cells, and are involved in cell-to-cell signaling in development and tissue homeostasis; how this sophisticated sensory system arose has been little-studied and its evolution is key to understanding how sensation arose in the Animal Kingdom. Sponges (Porifera), one of the earliest evolving phyla, lack conventional muscles and nerves and yet sense and respond to changes in their fluid environment. Here we demonstrate the presence of non-motile cilia in sponges and studied their role as flow sensors.
Results
Demosponges excrete wastes from their body with a stereotypic series of whole- body contractions using a structure called the osculum to regulate the water- flow through the body. In this study we show that short cilia line the inner epithelium of the sponge osculum. Ultrastructure of the cilia shows an absence of a central pair of microtubules and high speed imaging shows they are non- motile, suggesting they are not involved in generating flow. In other animals non-motile, primary, cilia are involved in sensation. Here we show that molecules known to block cationic ion channels in primary cilia and which inhibit sensory function in other organisms reduce or eliminate sponge contractions. Removal of the cilia using chloral hydrate, or removal of the whole osculum, also stops the contractions; in all instances the effect is reversible, suggesting that the cilia are involved in sensation. An analysis of sponge transcriptomes shows the presence of several transient receptor potential (TRP) channels including PKD channels known to be involved in sensing changes in flow in other animals. Together these data suggest that cilia in sponge oscula are involved in flow sensation and coordination of simple behaviour.
Conclusions
This is the first evidence of arrays of non-motile cilia in sponge oscula. Our findings provide support for the hypothesis that the cilia are sensory, and if true, the osculum may be considered a sensory organ that is used to coordinate whole animal responses in sponges. Arrays of primary cilia like these could represent the first step in the evolution of sensory and coordination systems in metazoans.
In a separate paper - ‘The analysis of eight transcriptomes from all Porifera classes reveals surprising genetic complexity in sponges’, her lab found -
Our analyses showed that all sponge classes share an unexpectedly large complement of genes with other metazoans. Interestingly, hexactinellid, calcareous and homoscleromorph sponges share more genes with bilaterians than with non-bilaterian metazoans. We were surprised to find representatives of most molecules involved in cell-cell communication, signaling, complex epithelia, immune recognition and germ-lineage/sex, with only a few, but potentially key, absences.
All those exciting results are summarized in her recent review - ‘Elements of a ‘nervous system’ in sponges’.
-——————————————–
Casey Dunn and siphonophores

Casey Dunn’s lab focuses on siphonophores. If you do not know anything about those strange animals, please take a look at this beautiful site prepared by him. In his research, he combines biological methods and bioinformatics to study these strange creatures, and we mentioned some of his papers in the evolutionary biology section. Curious readers may start from his 2008 Nature paper and continue to two recent ones uploaded in biorxiv.
The Histology of Nanomia bijuga (Hydrozoa: Siphonophora)
The siphonophore Nanomia bijuga is a pelagic hydrozoan (Cnidaria) with complex morphological organization. Each siphonophore is made up of many asexually produced, genetically identical zooids that are functionally specialized and morphologically distinct. These zooids predominantly arise by budding in two growth zones, and are arranged in precise patterns. This study describes the cellular anatomy of several zooid types as well as of the stem and gas-filled float, called the pneumatophore. The distribution of cellular morphologies across zooid types enhances our understanding of zooid function. The unique absorptive cells in the palpon, for example, indicate specialized intracellular digestive processing in this zooid type. Though cnidarians are usually thought of as mono-epithelial, we characterize at least two cellular populations in this species which are not connected to a basement membrane. This work provides a greater understanding of epithelial diversity within the cnidarians, and will be a foundation for future studies on Nanomia bijuga, including functional assays and gene expression analyses.
Stem cells in a colonial animal with localized growth zones
Siphonophores (Hydrozoa) have unparalleled colony-level complexity, precision of organization, and functional specialization between zooids (i.e., the units that make up colonies). Previous work has shown that, unlike other colonial animals, most growth in siphonophores is restricted to one or two well-defined growth zones that are the sites of both elongation and zooid budding. To understand this unique growth at the cellular level, we characterize the distribution of interstitial stem cells (i-cells) in the siphonophore Nanomia bijuga. Within the colony we find that i-cells are present at the tips of the growth zones, at well-defined sites where new zooid buds will arise, and in the youngest zooid buds. As each zooid matures, i-cells become progressively restricted to specific regions until they are mostly absent from the oldest zooids. We find no evidence of the migratory i-cells that have been observed in colonial cnidarian relatives. The restriction of i-cells to particular developing structures and sites of growth suggest a plant-like model of growth for siphonophores, where the growth zones function much like meristems. This spatial restriction of stem cells could also explain the precision of colony-level organization in siphonophores as a consequence of restricted growth potential.
-——————————————–
Is animal evolution gain in complexity or rearrangement of existing complexity?
All these exciting work may completely overhaul our understanding of evolution. If sponges and jellies have the same genetic toolkit as humans and functions like ‘nervous system’, is animal evolution rearrangement of existing complexity prepackaged in unicellular protists, such as choanoflagelletes? Does that mean the complexity came primarily from the unexplained evolution of eukaryote from prokaryote? That is puzzling, because the prokaryote–>eukaryote evolution is seen as saltatory based on evidences collected so far. Hopefully, recent technological advances will help us recover various missing blocks in fundamental understanding of evolution from prokaryote–>eukaryote.