
Comments on Our New Paper - "Sequence Analysis and Comparative Study of the Protein Subunits of Archaeal RNase P"
How did life originate on earth? What chemical properties of living objects make them different from the non-living objects? Where did the genome come from in the first place? How was life before the emergence of the genetic code? Why does the genetic code have that specific form? What biochemical advantages do the proteins get by having methionine as their first amino acid? How did the metabolic pathway evolve to its current form? How did bacteria, archaea and eukaryotes evolve into distinct kingdoms? How did multi- cellularity evolve? How did our ability to hear and smell evolve? Where did the adaptive immune system come from?
There are hundreds of such unanswered questions in biology and biochemistry. In fact, if one opens an introductory biology or biochemistry book and places ‘why’ in front of every statement, one may be surprised by the lack of available answers for most. Many ‘accepted’ answers to evolutionary questions are no more sophisticated than the Kipling’s Just So Stories after the technical mumbo-jumbos are cleaned away. Even the molecular answers are often derived from just one or two organisms (E. coli bacteria and S. cerevisiae eukaryote), and may not survive in the same form, when a third kingdom (archaea) is added to the mix.
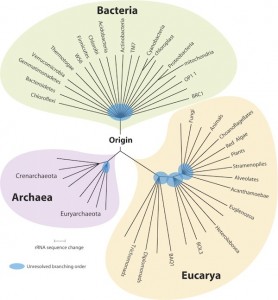
The availability of fully sequenced genomes may provide clues to solve many of those problems, and therefore scientists with computational skills looking for interesting puzzles will find a gold-mine (or many) in those unexplored questions. That realization is what got me attracted to this new line of research. All you have to do is get the ~50,000 bacterial genomes, ~6,000 archaeal genomes and >500 (unicellular) eukaryotic genomes sequenced so far, and then ask whether the accepted answer for a question is valid or not. In fact, I can make the first part easy for those, who are interested, because I am keeping an updated collection of those genomes (plus annotations) for my own work.
-———————————
With that general introduction, let me write more about the current paper. The abstract is presented below, and instead of copying the technical details from the paper, I will discuss some of the conceptual thinking that went into choosing this particular topic.
RNase P, a ribozyme-based ribonucleoprotein (RNP) complex that catalyzes tRNA 5?-maturation, is ubiquitous in all domains of life, but the evolution of its protein components (RNase P proteins, RPPs) is not well understood. Archaeal RPPs may provide clues on how the complex evolved from an ancient ribozyme to an RNP with multiple archaeal and eukaryotic (homologous) RPPs, which are unrelated to the single bacterial RPP. Here, we analyzed the sequence and structure of archaeal RPPs from over 600 available genomes. All five RPPs are found in eight archaeal phyla, suggesting that these RPPs arose early in archaeal evolutionary history. The putative ancestral genomic loci of archaeal RPPs include genes encoding several members of ribosome, exosome, and proteasome complexes, which may indicate coevolution/coordinate regulation of RNase P with other core cellular machineries. Despite being ancient, RPPs generally lack sequence conservation compared to other universal proteins. By analyzing the relative frequency of residues at every position in the context of the high-resolution structures of each of the RPPs (either alone or as functional binary complexes), we suggest residues for mutational analysis that may help uncover structure-function relationships in RPPs.
-——————————-
Connecting genetics with physical/chemical laws
There is a physical and chemical underlayer in all living organisms, on top of which the genetic layer stays active. That is self-evident given that the living organisms have to satisfy all known laws of physics and chemistry, or have to have additional laws. The discovery of later will be news by itself.
Sadly, very few researchers with expertise in bioinformatics are looking into the connection between genomic information and the physical/chemical system. I came to the realization after working on the electric eel genome project, where we tried to figure out the genetic and evolutionary basis of electric eels giving massive electric shocks. In fact, those non-traditional organisms are so little explored that researchers can make unusual discoveries without the help from millions of dollars of grants. For example, check this single author paper with simple experiments that led to a profound discovery - “The shocking predatory strike of the electric eel”.
Just like electric eels can help in connecting electricity and genetics, RNAs like tRNAs and RNase P can connect the knowledge of chemistry with genetics and provide an evolutionary basis. That is the primary reason for my interest in RNase P.
-——————————-
Why RNase P and tRNA?
In my very personal opinion, tRNA and RNase P are the molecules, which connect the information content of the genome and the chemical underlayer of the cells. Moreover, recent discoveries suggest that at least one of these molecules (tRNA) is much less understood than previously thought. For example, the paper “The structural basis for specific decoding of AUA by isoleucine tRNA on the ribosome” by Rebecca M. Voorhees (well-respected senior authors Uttam RajBhandary and V Ramakrishnan) concludes -
The role of the agmatidine and lysidine modifications in isoleucine decoding is a notable example of the essential function of modifications in tRNA and, more generally, in RNA biology. It is increasingly evident that although the genetic code was first elucidated over 40 years ago, we are just beginning to appreciate the sheer complexity of ensuring accuracy in protein synthesis on the ribosome.
RNase P, whose function is intricately linked to tRNA is even less studied with the aid of so many sequenced genomes than tRNA. RNase P is a ribonucleoprotein complex, whose RNA component is present as ribozyme in bacteria, archaea and eukaryote. Given that the enzyme performs the mundane task of cutting 5’ end of precursor tRNA, why did it not get replaced by a protein early in the course of evolution? On the other hand, if the RNA has undiscovered major roles to play, why is it present in exactly one copy in all bacterial and archaeal genomes, and two (diverged) copies in eukaryotic genomes? Questions like these got me curious about tRNA and RNase P.
-——————————-
Why archaea?
Imagine you are a naturalist from 18th century looking for previously unknown large animals, and you are aware of four disconnected masses of land (Eurasia, Africa, Americas and Australia) with the first three well-explored. Where will you look for novelty? In the context of life on earth, archaea represents the unexplored continent.
-——————————-
The paper
Based on available genomes sequences and structures, we looked into the evolution of protein components of RNase P in archaea. It is known that the evolutionary origin of the protein component of the complex is quite puzzling. Its RNA component likely emerged before the origin of genetic code or all the way back in the ‘RNA world’, but the protein component appeared in an unusual manner. It has only one member in bacteria, five in archaea and nine or more in eukaryota. The single bacterial protein component has no match to anything in archaea or eukaryota, although all protein components in archaea have orthologs in eukaryotes.
In the paper, we searched the entire collection of sequenced archaeal genomes for RNase P proteins, checked their genomic neighborhoods, analyzed phylogeny and compared the conserved amino acid locations with protein structures. One unexpected observation was that all five proteins were present in all phyla of archaea. We expected them to get added to progressively to various phyla, and thought we could rank archaeal evolution based on the timing of addition. That did not happen opening up a hole in the general understanding of evolution. Did the entire complex in archaea appear all together very early during archaeal evolution, while the bacterial side got an entirely different protein? Proteins related to DNA replication and few other major cellular tasks also show similar behavior. That means there was a period after separation of bacteria and archaea, but before the separation of major archaeal phyla, when many important components of archaea appeared. How long was that period, and how did molecular evolution take place in that era?
Some of evolutionary questions related to RNase P can be addressed through better understanding of the functionality of the complex, and that is where our sequence-structure comparison will come in handy. We did all necessary computational analysis to help an experimentalist initiate mutagenesis on the highly conserved locations of the proteins and understand how the amino acids work together with the ribozyme RNA to cut the tRNA.
-——————————-
Collaborators


The most enjoyable aspect of this project was my collaborators (Professor Venkat Gopalan and his student Stella Lai, and Professor Charles Daniels - all from Ohio State University). Each of us brought unique expertise to the project. Dr. Gopalan’s lab specializes on RNase P, whereas Dr. Daniels is an expert on archaea and especially halobacteria. If there is one big take home message I took from this project, it is that working with small groups asking focused questions is always a lot more exciting than ‘big science’.