
Improved the Search Routine in Our Website
We greatly improved the search function at the top of the page to bring the world of transcriptome closer to you.

Clicking on the link takes you to a different page, where you can search through all transcriptome experiments done by researchers around the globe. So far, we incorporated information from NCBI GEO, NCBI SRA and EBI Arrayexpress repositories.
If you type a database ID in the search box (such as ‘GPL198’ from GEO or ‘E-AFMX-1’ from Arrayexpress), whole lot of information on that specific experiment are displayed. If you search with regular words, we try to list many relevant experiments from different databases, and then clicking on the corresponding links gives you more details on the experiments.
There are many creative ways to use the search tool. Let us give one example. You probably remember from our trends section that GPL570 is the transcriptome platform most popular among researchers. It is an oligonucleotide array from Affymetrix with probes from all human genes.
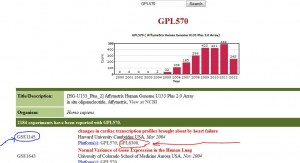
Suppose you like to use GPL570 for your research, and are looking for creative ideas. For example, many researchers combine some other array(s) with GPL570 to enhance their experiments. If you type GPL570 in the search box, you find what the platform GPL570 consists of, all reported experiments using the array (GSE1145 and GSE1643 in the following figure) and also all associated arrays/platforms used along with GPL570 by various researchers (e.g. GPL8300 in the following figure). Clicking on each link gives more information on the experiment or platform.
For those trying to analyze data or compare their data with published studies, we are planning to display as much useful information as possible. So far, we included information from all manually curated sets at NCBI. For example, if you type ‘GDS1001’ in the search box, it will let you explore the experiment GSE1134 and find up/downregulated genes at NCBI. We are expecting to add links to all available tools, as well as install few of our own.
I hope you find the new features useful. Please comment and let us know.