
Contrail - A de Bruijn Genome Assembler that uses Hadoop
Today’s commentary will tie three of our themes together - (i) de Bruijn graph-based genome or transcriptome assemblers, (ii) is there any alternative to buying a computer with huge RAM, and (iii) Hadoop. The question in (ii) occurs frequently in the mind of any bioinformatician or manager of core facility working on NGS sequences, because RAMs are very expensive. Hadoop is a good alternative used to process petabytes of data by internet companies (yes, they talk about petabyte scale, no kidding), but there is some barrier to entry for non-practitioners in using it. I promise that I won’t leave you high and dry here with some abstract discussion. By the end of this post, you will be able to assemble a (small) sequence library using a Hadoop-based approach and more or less understand what is being done by the assembler. The approach is scalable. If you find a large Hadoop setup at your computer cluster or Amazon elastic Mapreduce, you can play on your large data sets using the same commands.
We shall use a de Bruijn graph-based genome assembler named Contrail. It is developed by Michael Schatz at Cold Springs Harbor lab, who worked closely with the leading group of researchers at Center for Bioinformatics and Computational Biology at University of Maryland. This poster explains how the program works, and you will find many familiar themes already covered by our de Bruijn graph series. The program is available for download at sourceforge, but I did not find any detail on how to install and run it. That should not deter advanced users like you and me in using it. Here is a short manual I developed for using the program.
Step 1.
Go to the sourceforge svn link for Contrail and download gnu tarball for the code. Uncompress the files, and you will see the following directory structure (click on the image to see a larger view).
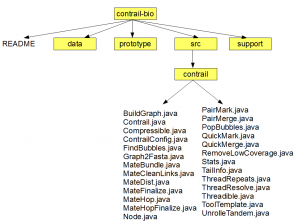
The directory structure is correct, but I did not show detail contents of all directories except one, where the source is located.
Step 2.
If you open any of the Java programs in the ‘contrail-bio/src/contrail’ directory, you will find that they are written in the same format as described in our Hadoop example. Each file has some headers at the top, a Mapper class, a Reducer class and a main program. You will see some small differences too. The ‘main’ program invokes a run routine, which has all the details about invoking map, reduce, etc., whereas, in our example, we placed all those codes in the ‘main’ program itself. Another difference you will see is that the format of Mapper, Reducer, etc. are following the depraced Wordcount example written for older version of Hadoop, and not the latest version of Hadoop used by us. Check our FAQ section for more details about the difference. Using older code will generate some annoying warning at the compilation time, but the program will run fine.
I mentioned the code for getting you feel comfortable about how the assembler works, but we do not really need to understand it for what we plan to do here. You are already familiar about how to compile Hadoop MapReduce Java programs from our previous example. Let’s get our hands dirty with Contrail.
Assuming that you have hadoop-0.20.2 installed, we will compile the Java programs in ‘contrail-bio/src/contrail’ directory using the following procedure. It is very similar to the example that you worked out.
% cd contrail-bio/src/contrail
% mkdir contrail_classes
% javac -Xlint:all -Xlint:deprecation -Xlint:unchecked -classpath ../../../hadoop-0.20.2/hadoop-0.20.2-core.jar:../../../hadoop-0.20.2/lib /commons-cli-1.2.jar:../../../hadoop-0.20.2/lib/log4j-1.2.15.jar -d contrail_classes BuildGraph.java Compressible.java Contrail.java ContrailConfig.java FindBubbles.java Graph2Fasta.java MateBundle.java MateCleanLinks.java MateDist.java MateFinalize.java MateHop.java MateHopFinalize.java Node.java PairMark.java PairMerge.java PopBubbles.java QuickMark.java QuickMerge.java RemoveLowCoverage.java RemoveTips.java Stats.java TailInfo.java ThreadRepeats.java ThreadResolve.java Threadible.java ToolTemplate.java UnrollTandem.java
% jar -cvf contrail.jar -C contrail_classes .
The syntax is very similar to before except the following changes.
i) “-Xlint:all -Xlint:deprecation -Xlint:unchecked” is added to reduce the number of ‘deprecated code’ warnings, because the code is not written for latest hadoop we are compiling it in.
ii) We added an additional jar library -
"../../../hadoop-0.20.2/lib/log4j-1.2.15.jar".
iii) Instead of compiling only one Java program in the previous Kmer example, we are compiling all Java programs together.
Irrespective of the differences, at the end of the day you get one jar file (contrail.jar) to be used with Hadoop.
Next, we will run this jar file under Hadoop to assemble some reads present in ‘contrail-bio/data’ directory. The method is identical to the Kmer example. Go to ‘contrail-bio’ directory and run -
% ../hadoop-0.20.2/bin/hadoop jar src/contrail/contrail.jar contrail.Contrail
ERROR: -asm is required
ERROR: -k is required
ERROR: -reads is required
The program is telling you that you have to give those three inputs. Let us
run it properly.
% mkdir assembly
% ../hadoop-0.20.2/bin/hadoop jar src/contrail/contrail.jar contrail.Contrail
-asm assembly -k 25 -reads reads data
The program will run for a few minutes generating all kinds of Hadoop-related messages. Once complete, you will see the assembly directory full with various directories. The assembly you are interested in is located in ‘assembly/99-final.fa’ directory.
If you want to run this program on a large Hadoop cluster, I would recommend you to do it in two steps. First, try the Kmer example that I gave you in previous example. That is the simplest possible bioinformatics program on NGS sequences using Hadoop. If that runs fine, you will know that your cluster setup, size, etc. are fine and you are ready to play big league.
In the next posts of this series, we will look into changing other parameters of Contrail program and try to understand the algorithm.