
What is the Cost of Scaling Up Genome Assembly Massively?
@zaminiqbal posted the link to an interesting genome assembly program called “SWAP-Assembler” presented as a scalable and fully parallelized genome assembler.
There is a growing gap between the output of new generation massively parallel sequencing machines and the ability to process and analyze the sequencing data. We present SWAP-Assembler, a scalable and fully parallelized genome assembler designed for massive sequencing data. Intend of using traditional de Bruijn Graph, SWAP-Assembler adopts multi-step bi-directed graph (MSG). With MSG, the standard genome assembly (SGA) is equivalent to the edge merging operations in a semi-group. Then a computation model, SWAP, is designed to parallelize semi-group computation. Experimental results showed that SWAP-Assembler is the fastest and most efficient assemblers ever, it can generated contigs with highest accuracy over all five selected assemblers and longest contig N50 in all selected parallel assemblers. Specially, in the scalability test, SWAP-Assembler can scales up to 1024 cores when processing Fish and Yanhuang dataset, and finishes the assembly work in only 15 and 29 minutes respecitively.
The associated paper was published last year -
Improved Parallel Processing of Massive De Bruijn Graph for Genome Assembly
Finishing human genome assembly in 30 minutes is incredibly fast, but at what cost? We asked in twitter and several people helped us out.
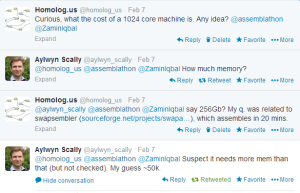
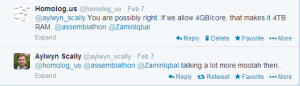
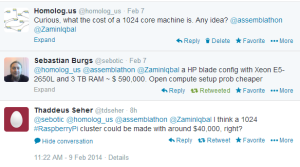
It is important to remember that Raspberry Pi contains only 256MB to 512MB, in case memory is a constraint for Swap-assembler.