
Posters and Slides from #BoG14
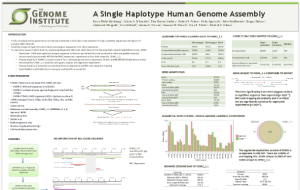
1. K. M. Steinberg - A Single Haplotype Genome Assembly (Please click on image to view slide at figshare)
2. Christopher DeBoever - SF3B1 mutations in different cancer types cause recognition of sterically hindered cryptic splice sites downstream of the branch point

3. Jia-Xing Yue - Insight into cephalochordate evolution from a genomic study of the Bahama amphioxus, Asymmetron lucayanum (changed mind and requested us to take down, because of unpublished data)
Cephalochordates (commonly known as amphioxus or lancelets) refers to a group of marine animals that are the modern survivors of one ancestral chordate lineage whose phylogenetic position falls at the boundary between invertebrates and vertebrates. Previous study showed that they possess many primitive features in morphology, development and also genomic architecture and thus are considered as the best living proxy for ancestral chordates before the rise of vertebrates. In this study, we performed both RNA-Seq and whole genome shotgun sequencing for the Bahama amphioxus, Asymmetron lucayanum. The comparison between Asymmetron lucayanum with another distantly related cephalochordate species, Branchiostoma floridae, together with several representative vertebrate species was conducted for both coding and noncoding regions. For the coding regions, we found cephalochordates evolve more slowly than vertebrates, which is consistent with their apparent morphological evolutionary stasis. However, genes involved in innate immunity are notably evolving faster. For the noncoding regions, we identified ~170,000 cephalochordate conserved noncoding elements (cCNEs) shared between Asymmetron and Branchiostoma using on a reads-mapping based approach. We also examined the genomic distribution, functional association and evolutionary trajectories of these cephalochordate CNEs in the context of chordate evolution.
4. Unknown author, who does not lack originality -
