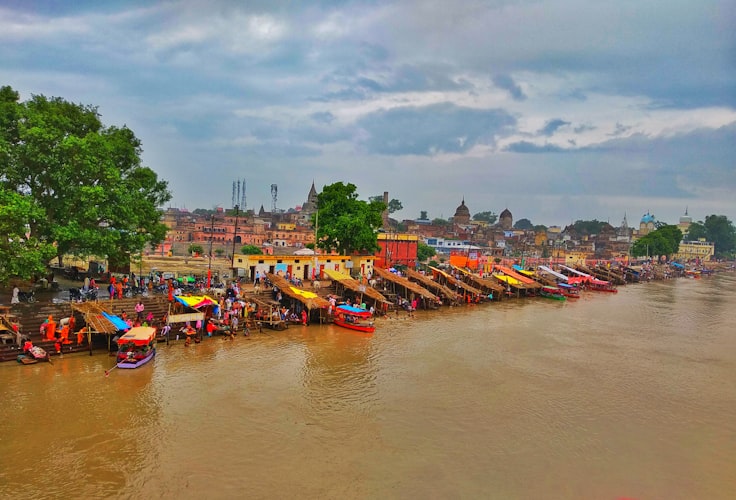

MEGAHIT - An Ultra-fast Single-node Solution for Large and Complex Metagenomics Assembly via Succinct de Bruijn Graph
About two years back, we reported about Succinct de Bruijn Graph construction by Alex Bowe and collaborators. Also, earlier this year, HKU group of professor Tak-Wah Lam published their implementation of GPU-Accelerated BWT Construction for Large Collection of Short Reads. Now those two are combined along with ideas from IDBA-UD into a metagenome assembler. The paper is available from arxiv.
MEGAHIT is a NGS de novo assembler for assembling large and complex metagenomics data in a time- and cost-efficient manner. It finished assembling a soil metagenomics dataset with 252Gbps in 44.1 hours and 99.6 hours on a single computing node with and without a GPU, respectively. MEGAHIT assembles the data as a whole, i.e., it avoids pre-processing like partitioning and normalization, which might compromise on result integrity. MEGAHIT generates 3 times larger assembly, with longer contig N50 and average contig length than the previous assembly. 55.8% of the reads were aligned to the assembly, which is 4 times higher than the previous. The source code of MEGAHIT is freely available at this https URL under GPLv3 license.