
Top Ten Genomes - (i) Lamprey
The Biology of Genomes conference is taking place at CSHL this week, and therefore our blog will talk a lot about genomes and researchers who work on genomes. After going through many genome papers published over the last decade and more, we decided to pick a set of ten most interesting genomes. Please feel free to suggest your choices, if you disagree.
-————————
Our award for the most interesting genome goes to lamprey.
1. Evolutionary position
For researchers interested in studying vertebrate biology, the lampreys hold a special position in the phylogeny tree. The following figure from an UCL website explains why.
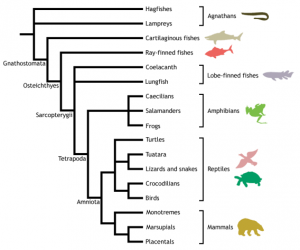
Lampreys dominated the seas before vertebrates with jaws came into existence, and their genomes can tell us about the evolution of jawed vertebrates. Lack of jaw does not mean lampreys go hungry in competition with other fish. The following video can tell you how they live well by sucking blood from other animals.
This blood-sucking property of Lamprey had been noticed by people since ancient times, including an unusually cruel Roman ruler.
Publius Vedius Pollio (died 15 BC) was a Roman equestrian of the 1st century BC, and a friend of the Roman emperor Augustus, who appointed him to a position of authority in the province of Asia. In later life he became known for his luxurious tastes and cruelty to his slaves when they displeased him, he had them fed to lampreys (or morays) that he maintained for that purpose, which was deemed to be an exceedingly cruel act. When Vedius tried to apply this method of execution to a slave who broke a crystal cup, Emperor Augustus (Pollio’s guest at the time) was so appalled that he not only intervened to prevent the execution but had all of Pollio’s valuable drinking vessels deliberately broken. This incident, along with Augustus’s demolition of the massive villa he inherited after Vedius’s death in 15 BC, were frequently referred to in antiquity in discussions of ethics and of the public role of Augustus.
-——————————-
2. VLR immune system
Lamprey genome does not only tell us about the evolution of jaws, but it also informs about the evolution of our adaptive immune system. In a remarkable discovery, Zev Pancer and colleagues found out about the presence of an entirely new adaptive immune system in lamprey.

Somatic diversification of variable lymphocyte receptors in the agnathan sea lamprey
Although jawless vertebrates are apparently capable of adaptive immune responses, they have not been found to possess the recombinatorial antigen receptors shared by all jawed vertebrates. Our search for the phylogenetic roots of adaptive immunity in the lamprey has instead identified a new type of variable lymphocyte receptors (VLRs) composed of highly diverse leucine-rich repeats (LRR) sandwiched between amino- and carboxy-terminal LRRs. An invariant stalk region tethers the VLRs to the cell surface by means of a glycosyl-phosphatidyl-inositol anchor. To generate rearranged VLR genes of the diversity necessary for an anticipatory immune system, the single lamprey VLR locus contains a large bank of diverse LRR cassettes, available for insertion into an incomplete germline VLR gene. Individual lymphocytes express a uniquely rearranged VLR gene in monoallelic fashion. Different evolutionary strategies were thus used to generate highly diverse lymphocyte receptors through rearrangement of LRR modules in agnathans (jawless fish) and of immunoglobulin gene segments in gnathostomes (jawed vertebrates).
Pancer and co-authors made their discovery without help from the genome sequence. When the lamprey genome was sequenced, it was found that the genome included orthologs of other components of vertebrate adaptive immune systems. Therefore, the immune system of lamprey is not at all ‘primitive’.
Lamprey immunity is far from primitive
The adaptive immune systems of vertebrates provide remarkable examples of evolutionary innovation. This is most evident in the unusual mechanisms that jawed vertebrates have invented to create and deploy T-cell receptor and Ig diversity. In a series of papers published over the past decade (detailed below), the variable lymphocyte receptors of the jawless vertebrates (VLRs) (Fig. 1A) have emerged as an equally powerful example of evolutionary novelty. This story of parallel solutions to the challenge of nonself recognition is made all of the more compelling when one considers the wider system in which these receptors function. Although the V(D)J system of mammals and the VLR system of the agnathans are structurally unrelated, their diversity is expressed and selected in the context of lymphocytic cells that at some level share homology. This similarity offers enormous opportunity for understanding how novel immune mechanisms evolve and are incorporated into a background of more ancient cell regulatory networks. In PNAS, Das et al. (1) present another advance in the understanding of the VLR-based adaptive immune system. Using the lamprey genome sequence (2) and cDNA sequences, the authors characterize the generation of diversity in the third lamprey VLR locus (VLRC) and establish yet another variation on how immune specificity is created.
Given that the Biology of Genome conference always includes those making major genomic discoveries as keynote speakers, Zeev Pancer could have easily become the one in 2015 for his remarkable discovery. Sadly he passed away in 2014 at a relatively young age (56).
-——————————-
3. Genome shrinks

In another unusual observation about the lamprey genome, Jeramiah Smith from Chris Amemiya’s group reported that a large chunk (~20%) got deleted, when the lampreys developed from single cell to multi-cellular organism. That is very surprising and rather unexpected for a genome. How does it know, which parts to get rid of? Nobody knows that answer yet. In fact, it was observed that the deleted parts included some coding genes and not just junk DNA.
Fish Throws Away Its Genes as It Grows
Whether it’s its extraterrestrial looks or status as a “living fossil,” there’s always been something fishy about the sea lamprey. Now scientists have added another oddity to the creature’s repertoire: The lamprey jettisons 20% of its genome during development.
Jeramiah Smith of the University of Washington, Seattle, first suspected something strange while piecing together the sea lamprey’s genetic sequence. The postdoctoral fellow and his colleagues tried labeling live lamprey cells using a technique that detects broken DNA. “Every cell in the embryo was [labeled] as dying,” he recalls. So he took a closer look to see what was going on and got a big surprise.
Working with Chris Amemiya of the Benaroya Research Institute at Virginia Mason in Seattle, the group found that lamprey sperm DNA had sequences not found in lamprey liver and that overall the sperm genome was millions of bases longer. The sperm DNA included a highly repetitive sequence called Germ1, and by monitoring the loss of Germ1 during development, Smith was able to track the genome’s reorganization.
-——————————-
4. Genome assembly

[Photo courtesy: Chris Amemiya]
Those working on genome assembly also finds the lamprey genome a rewarding challenge. Regions of this GC-rich genome are inaccessible to sequencing technologies with GC-bias. To add to the troubles, the genome is highly polymorphic and very hard to assemble. To add to the troubles, the genome has nearly 100 pairs of (nuclear) chromosomes !!
The somatic genome of sea lamprey was published in 2013.
Lampreys are representatives of an ancient vertebrate lineage that diverged from our own ~500 million years ago. By virtue of this deeply shared ancestry, the sea lamprey (P. marinus) genome is uniquely poised to provide insight into the ancestry of vertebrate genomes and the underlying principles of vertebrate biology. Here, we present the first lamprey whole-genome sequence and assembly. We note challenges faced owing to its high content of repetitive elements and GC bases, as well as the absence of broad-scale sequence information from closely related species. Analyses of the assembly indicate that two whole-genome duplications likely occurred before the divergence of ancestral lamprey and gnathostome lineages. Moreover, the results help define key evolutionary events within vertebrate lineages, including the origin of myelin-associated proteins and the development of appendages. The lamprey genome provides an important resource for reconstructing vertebrate origins and the evolutionary events that have shaped the genomes of extant organisms.
-——————————-
5. Six hox clusters

In another surprising observation, Tarang Mehta from B. Venkatesh’s group in Singapore discovered the lamprey genome to have six Hox clusters. Why is that unusual? The number of Hox clusters is an indicator of the number of times the organism’s genome went through whole-genome duplication. Based on position on phylogenetic tree, it was believed that lampreys went through one or two rounds of whole genome duplication, and therefore have two or four hox clusters. The observation about six hox clusters complicates everything we know about vertebrate genome evolution and whole genome duplication.
Evidence for at least six Hox clusters in the Japanese lamprey (Lethenteron japonicum)
Lampreys and hagfishes (cyclostomes) are the only living group of jawless vertebrates and therefore are important for the study of vertebrate evolution. We have characterized Hox clusters in the Japanese lamprey (Lethenteron japonicum), and shown that it contains at least six Hox clusters as compared with four Hox clusters in tetrapods. This suggests that the lamprey lineage has undergone an additional round of genome duplication compared with tetrapods. Several conserved noncoding elements (CNEs) were predicted in the Hox clusters of lamprey, elephant shark, and human. Transgenic assay of CNEs demonstrated their potential to function as cis-regulatory elements. Thus, these CNEs may represent part of the core set of cis-regulatory elements that were present in the common ancestor of vertebrates.
-——————————–
6. Evolution of Vertebrate Neural Crest and Gene Regulatory Network
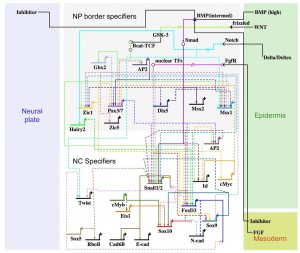
Eric Davidson’s latest book includes several sections on lamprey. That is no surprise given that lampreys hold an unique position in phylogenetic tree to connect sea urchins and vertebrates. Davidson’s lab focuses on sea urchin, and Marianne Bronner, one of his Caltech colleagues, is using Davidson’s gene regulatory network methodlogy to study the evolution of vertebrate neural crest, with lamprey being the organism of her lab’s focus.

-——————————————–
7. Life cycle of sea lamprey
Chris Amemiya added that the life history of sea lamprey is equally interesting. The following picture from a New York State Department website explains it well, and to find the unusual part, check the time periods marked near the periphery of the circle. These amazing creatures spend nearly four years as larvae and then only 12 months as adults !!
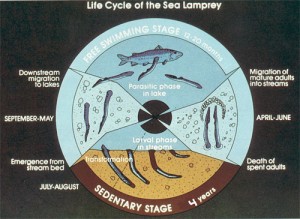
Life Cycle - How does it live and breed?
Ammocoetes
A Larval Lamprey
The blind worm-like larval lamprey, known as ammocoetes [am-mah-seats], can grow up to 5 inches long. They hatch from eggs in gravel nests in tributaries and drift downstream with the current. When they locate suitable habitat - usually silt/sand stream bottoms and banks in slower moving stretches of water
- they burrow in and take up residence, filter-feeding on algae, detritus and microscopic organisms and materials. In the Lake Champlain Basin this stage of the sea lamprey’s life cycle usually lasts 3 to 4 years; in other waters lamprey spend up to 10 years in their larval form.
Transformers
Sometime in mid to late summer of their third or fourth year the ammocoetes undergo a dramatic change in both form and function. They develop eyes and a suction disk mouth, and become a smaller version of the adult sea lamprey. Also during this stage their kidneys change to allow them to live in saltwater. Once the ammocoetes change is complete, the newly transformed sea lamprey, known as a transformer, leaves its burrow and moves downstream towards Lake Champlain. The sea lamprey is then ready to begin the next stage in its life as a parasite of fish. The juvenile sea lamprey move into deeper water and begin to seek host fish on which to feed.
Parasitic Juveniles
The juvenile sea lamprey uses its suction disk mouth which is filled with small sharp, rasping teeth and a file-like tongue to attach to fish, puncture the skin, and drain the fish’s body fluids. An anticoagulant in their saliva ensures that the blood of the host fish does not clot while the sea lamprey feed. Often the host fish die from loss of blood, or infections resulting from stress. Fish that survive sea lamprey attacks will have decreased reproduction. Sea lamprey in Lake Champlain prefer landlocked Atlantic salmon (salmon), lake trout and other trout species, due to their small scales and thin skin. The same native fish species prized by anglers, and that are such an important part of the natural ecosystem of the lake.
Sea lamprey also feed on other fish species, including lake whitefish, walleye, northern pike, burbot, and lake sturgeon. The lake sturgeon is listed as a threatened species in New York and an endangered species in Vermont and it is likely that sea lamprey are affecting their survival. Most sea lamprey hosts are native fish species that have been part of the Lake Champlain Basin ecosystem for thousands of years.
Spawning
In the spring, sexually mature adult sea lamprey migrate up tributaries to spawn. They locate spawning streams by following pheromones (naturally produced chemical attractants) released by ammocoetes living in those waters. A pair of male and female sea lamprey build a nest, called a redd, in a gravel stream bottom in section of flowing water. The female lays tens of thousands of eggs and the male fertilizes them, then having completed this act the sea lamprey die. The eggs lie in the small spaces between the gravel, and are provided oxygen by the flowing water. Weeks later the eggs hatch and the complex life cycle of the sea lamprey begins again.