
Pacbio Assembly and Phasing
Jason Chin, who wrote the HGAP assembler for PacBio-based genome assemble, is on to a more difficult problem. Here are his slides from the recent CSHL conference -
-———————————————–
Readers of our blog surely knows that a different way for doing these things was published last year. Please do not tell him :)
HapCompass: An Elegant Use of Graphs for Haplotype Assembly/Phasing
The novelty of HapCompass is to create a different kind of graph (named compass in the paper) that assigns SNPs as nodes and linking paired-end sequences as edges. In the following figure, if two SNPs come from the same chromosome, they will be linked by an edge. If not, no edge will exist between them. Of course, the previous statement assumes complete error-free world. Few spurious edges can be created by noisy sequence data.
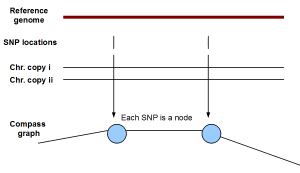
Based on the above graph structure, the authors translated the phasing/haplotype assembly problem into a spanning tree problem, and solved it through a minimum weight edge removal optimization algorithm.