
Bringing New Computing Hardware Architecture to 'Life'
Over the last few months, this blog covered bioinformatics programs and algorithms developed by leading groups around the world. Today we like to make an announcement about our own project that is interesting enough to deserve a place in this space.
We are working with Stillwater Supercomputing, Inc. and Pico Computing, Inc. to build a bioinformatic appliance with novel hardware architecture for accelerating bioinformatics algorithms. The first application of this technology will be on display at Supercomputing 12 in Salt Lake City, November 12-15.
With exponential increase in amount of sequence data from NGS technologies, access to computing power has become the biggest bottleneck. The problem manifests in two ways - (i) in lack of number of processors to rapidly analyze the data, (ii) shortage of memory (RAM, cache) to hold transient data that is being processed. If you think about both issues carefully, you will find them to be manifestations of inherent lack of parallelizability of Intel-based (von Neumann) computing architecture.
Researchers working on computing architectures realized the problems long back and proposed alternatives. The Knowledge Processing Unit, or KPU, patented by Stillwater Supercomputing, is a distributed data flow machine, a highly parallelizable architecture, that can seamlessly introduce thousand or more processing units in a chip. We leveraged this architecture to build a genome assembler and implement it on FPGA-based SC5 SuperCluster platform developed by Pico Computing, Inc.
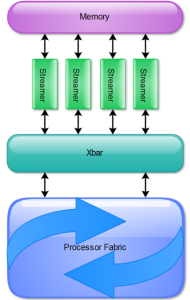
Access to Stillware hardware can be done by Stillwater Knowledge Processing Platform, a software abstraction layer, that solves the problem of hardware dependence for high-performance bioinformatics applications. As those types of bioinformatics applications may need to run on multi-core, many-core, GPUs and FPGAs, the desire is to decouple the bioinformatics algorithms from underlying parallel hardware. The Stillwater Knowledge Processing Platform provides this decoupling by creating a set of useful bioinformatics operators (k-mer counting, Smith-Waterman alignment, hashing, Bloom filter, etc.) and enables the application to run across wide spectrum of hardware without modifications. Furthermore, the Stillwater Platform can take advantage of multiple FPGA chips to create scalable supercomputing systems to match any required throughput for medium to large scale genome centers.
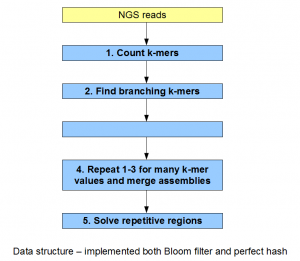
Our genome assembler implemented almost all assembly concepts discussed in our blog over the last few months. We were especially influenced by SPAdes, Minia, Meraculous and IDBA papers. The original purpose was to check which bioinformatic operators got used repeatedly, and then design their hardware-accelerated abstract forms. However, during the course of our work, the assembler project took a life of its own, and in the end, we believe we implemented a program that is cutting- edge by itself as a stand alone software.
Going forward, we like to leverage the hardware platform and software abstraction layer to parallelize various popular bioinformatics applications. Instead of reinventing the wheel in every case, we like to keep those programs in their original forms as much as possible, except for the part that gets accelerated.
Please come visit us at Pico Computing booth in Supercomputing 12 in Salt Lake City at Pico Computing booth to learn more about this exciting project.
Our regular commentaries will resume in the following week.