
Academic Bioinformaticians Uncomfortable with Illumina's Publication of Variant
The battle over cancer bioinformatics may be moving out of the hands of academic bioinformaticians. Last year, Broad announced changes in GATK license, irritating Sanger Institute so much that they dropped GATK altogether. That change made by Broad is not an one- off decision but rather a new ‘business model’. Their newly released Discovar variant assembler built on top of ALLPATHS-LG assembly code also contains the same license. In the meanwhile, BGI (Beijing Genomics ‘Institute’) acquired Complete Genomics, making academics wonder when Sanger Institute, Broad Institute and Systems Biology Institute will start gobbling up companies.
Today’s publication of a paper by Illumina did not make many bioinformaticians happy, if twitter comments are of any guide.
Isaac: Ultra-fast whole genome secondary analysis on Illumina sequencing plat forms
An ultrafast DNA sequence aligner (Isaac Genome Alignment Software) that takes advantage of high memory hardware (>48GB) and variant caller (Isaac Variant Caller) have been developed. We demonstrate that our combined pipeline (Isaac) is 4-5 times faster than BWA+GATK on equivalent hardware, with comparable accuracy as measured by trio conflict rates and sensitivity. We further show that Isaac is effective in the detection of disease-causing variants and can easily/economically be run on commodity hardware.
Availability: Isaac has an open source license and can be obtained at https://github.com/sequencing
Here are some twitter feedbacks from academics on the paper. The specifics about the paper may be right, but we do not agree with the general tone. Illumina’s Chesterford Research Park does have very good researchers, and Heng Li’s ropebwt is an implementation of an innovative algorithm developed by a group from there.
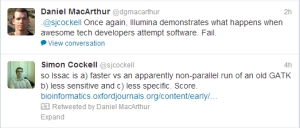
The real amateurs in Illumina are in their legal department. Based on our reading of various licenses coming out of Illumina, it is clear that those lawyers have any clue about how computer code is written.
The Isaac license is ‘Illumina open-source’, which makes the code free for non-commercial use. In other similar licenses, we looked hard to find out the implied meaning of commercial use, but thankfully Illumina makes the definition very clear.
Commercial Entity means for-profit entities. Academic universities and other bona fide non-profit entities are not Commercial Entities.
Commercial Useand grammatical variations (e.g., Commercial and Commercialize) means any use that is undertaken for the purpose of profit or other commercial gain, except use or sale of the Software as part of a SaaS Bundle, which is prohibited; see below. Sale of Software or products that include Software, use of Software in the provision of sequencing and other data-generating services (except not as a SaaS Bundle), and software support services are three examples (non-limiting) of Commercial Use.
………
If you would like to Commercialize the Software, including sell the Software, using it in services, or provide a service to support the Software, all in connection with non-Illumina branded equipment, then you are required to request and receive Illuminas permission before doing so.
What is the problem? Software program is not like a Illumina machine, and the real contribution is not in the packaged final product, but in the algorithms. True innovation is often in very small portions that defines the core algorithm, and such true innovations are hard to come by. So, if you release the code and put up a cumbersome license, smart people will learn the algorithm and implement the innovative part elsewhere. We will show an example with another of Illumina’s code that came with this cumbersome BEETL license. The result was this implementation of the same algorithm by Heng Li with improvements. Heng Li’s code is released under MIT license putting BEETL library out of business, when it comes to incorporation within other codes. Unfortunately, that will only have an impact on the lives of researchers like AJ Cox, MJ Bauer, T Jakobi T and G Rosone, while the lawyers will move on to prey on other hapless research groups.