Pacbio: Why We Stopped Using PacBioToCA and Lived Happily Thereafter
When we started working on PacBio data one year back, everyone recommended PacBioToCA. Pause for a moment to imagine how summer of 2012 was. Everyone was talking about Illumina, 454, de Bruijn graph, Velvet assembler and so on, and these ‘weird’ reads show up from nowhere. Using an analogy, everyone is talking about pizza and BioMickWatson shows five other foods that are like genome assembly, namely Eton mess, spaghetti Bolognese, Marmite, ‘macaroni’ cheese and anchovite. The initial impulse is to turn all those into toppings for pizza to make them attractive.
That is what PacBioToCA does. It turns PacBios into Illuminas and then let you forget about them. In detail, PacBioToCA painstakingly aligns all Illumina reads on to the PacBios and then locally assemble the Illumina reads. From that point onward, you are back to Illumina world. However, the alignment was incredibly time-consuming. LSC - the same story. It aligns all Illumina reads on to PacBio using Novoalign, an incredibly slow PacBio-unaware aligner. We realized that it made more sense to assemble Illumina reads first and then align them on PacBios.
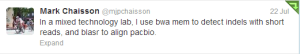
Over time we learned that any PacBio pipeline not using BLASR is not doing the analysis right. Mark Chaisson spent a lot of time to turn BLASR into an incredibly powerful tool. It includes the read-filtering program. It even has a PacBio read simulator, which, according to Mark, matches experimental data better than the published simulator PBSIM.
The main advantage of BLASR is its knowledge of indels being the primary mode of error in PacBio reads. So, it is very PacBio-aware, which other aligners are not.
-———————————–
Edit.
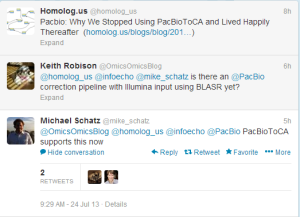
PacBioToCA also got upgraded, which we have not kept pace with. Here are the latest updates from Michael Schatz and others -
Also, Jason Chin mentioned a better approach -
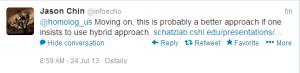
The linked presentations are available here and most possibly Jason is suggesting the following pipeline -
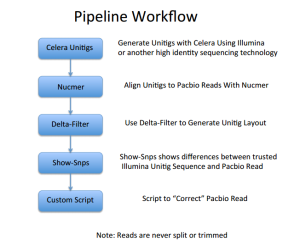

If you are using Mike Schatz’s method, the following twitter discussion may be of help to you.
@mike_schatz @aphillippy Is there any way to run mummer3/nucmer in multithread mode? I did some search but I can’t find related info.
– Jason Chin (@infoecho) July 25, 2013