
Sequencing miRNAs with NGS Technology
Mature miRNA sequences are 22 nucleotide long, and are perfect fit for next- generation sequencing technologies. As we explained in previous commentary, NGS technologies made the life of miRNA researchers very easy. In earlier days, miRNAs were primarily identified from genomes using computer programs, and there were not many genomes available. These days, researchers can do their own sequencing to study miRNA profiles in many different organisms, for which no genome is yet assembled.
Scientists studying miRNAs using NGS technologies face the opposite problem of those exploring protein-coding genes (RNAseq). Assembling the genes from short reads is a big challenge for RNAseq experiments. MiRNAs are at the other extreme being too short even for NGS. Therefore, typically a linker sequence is added to each miRNA to get the total length above 75-100 nt.
One advantage of using linker is the ability to multiplex many different tissues or samples. Usually, the throughput of HiSeq instruments is too high for sequencing one miRNA sample. So, combining many samples saves money. Multiplexing of different samples is done by adding different linkers to miRNAs in different samples.
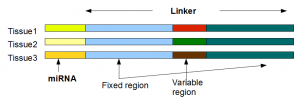
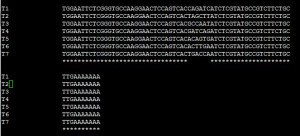
Those linkers are mostly the same with a small sample-specific variable part inserted in the middle. For illustration, we show the Clustal alignment for 7 linkers from a multiplexed miRNA profiling experiment in the next figure. The experiment used Illumina HiSeq sequencing technology with read length of 99 nucleotides. The linkers are about 74 nucleotide long with poly-A tails at the 3’ ends. Together the length of linker+miRNA is about 96 nucleotides.
In the next commentary, we will reverse engineer the output from one of those experiments to learn about some potential surprises.