Note: These tutorials are incomplete. More complete versions are being made available for our members. Sign up for free.

Impact of Changing k-mer Size
All previous examples considered k=7, but k can be any small or big integer. At its lowest limit, k can be 1. However, a k=1 de Bruijn graph is not very useful as you can see from the following figure.
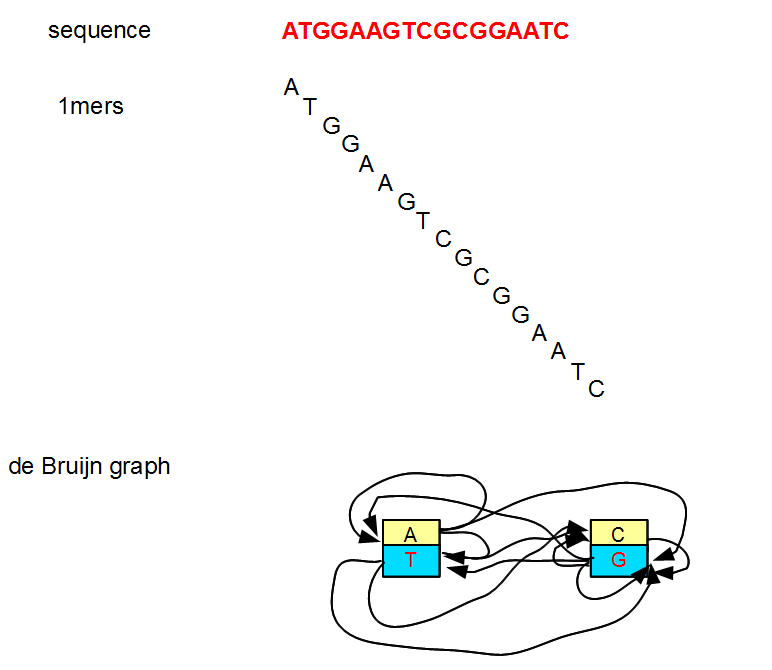
Genome assembly programs also avoid even k, because with even k, many k-mers become reverse complements of their own sequences. That causes ambiguities in the strand-specificness of the graph. Therefore, odd k-values are preferred.
De Bruijn assemblers like Velvet, Trinity, etc. use K-mers of odd length (21,23,25…). Here is the rationale. If K-mers are of even length, some K-mers can be reverse complements of themselves (e.g. ATATATATATAT). That will create ambiguity in the de Bruijn graph and make its resolution difficult. Palindromic K-mers can be always avoided with odd K-mer size, because the reverse complement of center nucleotide is different from the nucleotide itself.
In color space, reverse complement of a sequence is the reverse of the sequence. That means some odd-sized K-mers can be self-complementary, whereas even sized K-mers can never be so except for sequences like 111111111, 2222222, 3333333 and 44444444.
